Search
About Coprinellus micaceus str. FP101781 (GCA_004369175.1)
Coprinellus micaceus is a common species of mushroom-forming fungus in the family Psathyrellaceae with a cosmopolitan distribution. The fruit bodies of the saprobe typically grow in clusters on or near rotting hardwood tree stumps or underground tree roots. Depending on their stage of development, the tawny-brown mushroom caps may range in shape from oval to bell-shaped to convex, and reach diameters up to . The caps, marked with fine radial or linear grooves that extend nearly to the center, rest atop whitish stipes up to long. In young specimens, the entire cap surface is coated with a fine layer of reflective mica-like cells that provide the inspiration for both the mushroom's species name and the common names mica cap, shiny cap, and glistening inky cap. Although small and with thin flesh, the mushrooms are usually bountiful, as they typically grow in dense clusters. A few hours after collection, the gills will begin to slowly dissolve into a black, inky, spore-laden liquid—an enzymatic process called autodigestion or deliquescence. The fruit bodies are edible before the gills blacken and dissolve, and cooking will stop the autodigestion process. The microscopic characteristics and cytogenetics of C. micaceus are well known, and it has been used frequently as a model organism to study cell division and meiosis in Basidiomycetes. Chemical analysis of the fruit bodies has revealed the presence of antibacterial and enzyme-inhibiting compounds. Formerly known as Coprinus micaceus, the species was transferred to Coprinellus in 2001 as phylogenetic analyses provided the impetus for a reorganization of the many species formerly grouped together in the genus Coprinus. Based on external appearance, C. micaceus is virtually indistinguishable from C. truncorum, and it has been suggested that many reported collections of the former may be of the latter.
Taxonomy ID 71717
Data source DOE Joint Genome Institute
Comparative genomics
What can I find? Homologues, gene trees, and whole genome alignments across multiple species.
 More about comparative analyses
More about comparative analyses
 Phylogenetic overview of gene families
Phylogenetic overview of gene families
 Download alignments (EMF)
Download alignments (EMF)
Variation
This species currently has no variation database. However you can process your own variants using the Variant Effect Predictor:




 {#wiki_icon}
{#wiki_icon}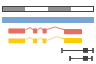
 Display your data in Ensembl Fungi
Display your data in Ensembl Fungi
 Update your old Ensembl IDs
Update your old Ensembl IDs
![Follow us on Twitter! [twitter logo]](/i/twitter.png)
