Search
About Morchella conica CCBAS932 (GCA_003790465.1)
Morchella conica is an old binomial name previously applied to species of fungi in the family Morchellaceae. It is one of three scientific names that had been commonly used to describe black morels, the others being M. angusticeps and M. elata. It was first introduced by mycologist Christian Hendrik Persoon in 1818, as a superfluous name for the old taxon Morchella continua. According to Richard and colleagues, Fries’ sanctioning applies only at the subgeneric level and the name is illegitimate. Throughout the years, the name M. conica has been invariably applied to many different species by different authors, and DNA analysis in 2014 revealed that morels identified as "M. conica" indeed belonged to Morchella deliciosa, Morchella purpurascens, Morchella tridentina, and Morchella vulgaris.
Taxonomy ID 1392247
Data source DOE Joint Genome Institute
Comparative genomics
What can I find? Homologues, gene trees, and whole genome alignments across multiple species.
 More about comparative analyses
More about comparative analyses
 Phylogenetic overview of gene families
Phylogenetic overview of gene families
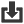 Download alignments (EMF)
Download alignments (EMF)
Variation
This species currently has no variation database. However you can process your own variants using the Variant Effect Predictor:




 {#wiki_icon}
{#wiki_icon}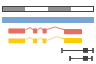
 Display your data in Ensembl Fungi
Display your data in Ensembl Fungi
 Update your old Ensembl IDs
Update your old Ensembl IDs
![Follow us on Twitter! [twitter logo]](/i/twitter.png)
