Search
About Ashbya gossypii ATCC 10895
Eremothecium gossypii (also known as Ashbya gossypii) is a filamentous fungus or mold closely related to yeast, but growing exclusively in a filamentous way. It was originally isolated from cotton as a pathogen causing stigmatomycosis by Ashby and Nowell in 1926. This disease affects the development of hair cells in cotton bolls and can be transmitted to citrus fruits, which thereupon dry out and collapse (dry rot disease). In the first part of the 20th century, E. gossypii and two other fungi causing stigmatomycosis (Eremothecium coryli, Aureobasidium pullulans) made it virtually impossible to grow cotton in certain regions of the subtropics, causing severe economical losses. Control of the spore-transmitting insects - cotton stainer (Dysdercus suturellus) and Antestiopsis (antestia bugs) - permitted full eradication of infections. E. gossypii was recognized as a natural overproducer of riboflavin (vitamin B2), which protects its spores against ultraviolet light. This made it an interesting organism for industries, where genetically modified strains are still used to produce this vitamin.
Picture credit: Public domain via Wikimedia Commons (Image source) Taxonomy ID 284811
(Text from Wikipedia.)
More information General information about this species can be found in Wikipedia
Taxonomy ID 284811
Data source Ashbya Genome Database
Comparative genomics
What can I find? Homologues, gene trees, and whole genome alignments across multiple species.
 More about comparative analyses
More about comparative analyses
 Phylogenetic overview of gene families
Phylogenetic overview of gene families
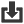 Download alignments (EMF)
Download alignments (EMF)
Variation
This species currently has no variation database. However you can process your own variants using the Variant Effect Predictor:
Regulation
What can I find? Microarray annotations.
Other Data
Probe mapping data has been loaded for the experiment A-AFFY-105.





 Display your data in Ensembl Fungi
Display your data in Ensembl Fungi
 Update your old Ensembl IDs
Update your old Ensembl IDs

![Follow us on Twitter! [twitter logo]](/i/twitter.png)
