Search
About Zygosaccharomyces bailii ISA1307 (GCA_000530735)
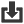
Zygosaccharomyces bailii is a species in the genus Zygosaccharomyces. It was initially described as Saccharomyces bailii by Lindner in 1895, but in 1983 it was reclassified as Zygosaccharomyces bailii in the work by Barnett et al.
Spoilage resulting from growth of the yeast Zygosaccharomyces is widespread, which has caused significant economic losses to the food industry. Within this genus, Z. bailii is one of the most troublesome species due to its exceptional tolerance to various stressful conditions. A wide range of acidic and/or high-sugar products such as fruit concentrates, wine, soft drinks, syrups, ketchup, mayonnaise, pickles, salad dressing, etc., are normally considered to be shelf-stable, i.e. they readily inactivate a broad range of food-borne microorganisms. However, these products are still susceptible to spoilage by Z. bailii.
(Text and image from Wikipedia, the free encyclopaedia.)
Taxonomy ID 1355161
Data source HMGU
Comparative genomics
What can I find? Homologues, gene trees, and whole genome alignments across multiple species.
 More about comparative analyses
More about comparative analyses
 Phylogenetic overview of gene families
Phylogenetic overview of gene families
 Download alignments (EMF)
Download alignments (EMF)
Variation
This species currently has no variation database. However you can process your own variants using the Variant Effect Predictor:




 Display your data in Ensembl Fungi
Display your data in Ensembl Fungi
 Update your old Ensembl IDs
Update your old Ensembl IDs
![Follow us on Twitter! [twitter logo]](/i/twitter.png)
