Search
About Aureobasidium pullulans (GCA_004917525.1)
Aureobasidium pullulans is a ubiquitous and generalistic black, yeast-like fungus that can be found in different environments (e.g. soil, water, air and limestone). It is well known as a naturally occurring epiphyte or endophyte of a wide range of plant species (e.g. apple, grape, cucumber, green beans, cabbage) without causing any symptoms of disease. A. pullulans can be cultivated on potato dextrose agar, where it produces smooth, faint pink, yeast-like colonies covered with a slimy mass of spores. Older colonies change to black due to chlamydospore production. Primary conidia are hyaline, smooth, ellipsoidal, one-celled, and variable in shape and size; secondary conidia are smaller. Conidiophores are undifferentiated, intercalary or terminal, or arising as short lateral branches. Endoconidia are produced in an intercalary cell and released into a neighboring empty cell. Hyphae are hyaline, smooth, and thinwalled, with transverse septa. The fungus grows at 10–35 °C with optimum growth at 30 °C. A. pullulans is notable for its phenotypic plasticity. Colony morphology may be affected by carbon source, colony age, temperature, light and substrate, with colonies ranging from homogeneous to sectored, yeast-like to filamentous growth, and from small to large. Besides these morphological plasticity A. pullulans is also adaptable to various stressful conditions: hypersaline, acidic and alkaline, cold, and oligotrophic. Therefore, it is considered to be polyextremotolerant. The genome also contains a homothallic mating-type locus. Despite the presence in the genome of a putative mating locus, and the observation of high recombination rates, no sexual cycle has yet been reported in this organism. Due to the relatively recent redefinition of the species, most published work does not yet distinguish between the new species belonging to the previously recognised A. pullulans species complex. It is therefore not clear to what extent this knowledge is valid for A. pullulans s. str. and what should be attributed to the three new species.
Taxonomy ID 5580
Data source Biotechnical Faculty, University of Ljubljana
Comparative genomics
What can I find? Homologues, gene trees, and whole genome alignments across multiple species.
 More about comparative analyses
More about comparative analyses
 Phylogenetic overview of gene families
Phylogenetic overview of gene families
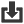 Download alignments (EMF)
Download alignments (EMF)
Variation
This species currently has no variation database. However you can process your own variants using the Variant Effect Predictor:




 {#wiki_icon}
{#wiki_icon}
 Display your data in Ensembl Fungi
Display your data in Ensembl Fungi
 Update your old Ensembl IDs
Update your old Ensembl IDs
![Follow us on Twitter! [twitter logo]](/i/twitter.png)
