Hortaea werneckii Assembly and Gene Annotation
About Hortaea werneckii (GCA_003704355.1)
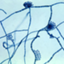
Hortaea werneckii is a species of yeast in the family Teratosphaeriaceae. Several salt-tolerance mechanisms of H. werneckii have been studied on molecular level. For example, it is known that its major compatible solutes are glycerol, erythritol, arabitol, and mannitol; melanin accumulation of the cell wall aids in retention of at least glycerol inside of the cell. Several components of the high osmolarity glycerol (HOG) signalling pathway (which controls responses to osmotic shock) have been studied in detail and some seem to differ in function compared to their counterparts in Saccharomyces cerevisiae. Whole genome sequencing of H. werneckii Genes encoding metal cation transporters, which are thought to play a role in halotolerance, experienced several additional gene duplications at various points during their evolution. A homothallic mating locus was found in all sequenced genomes, although one of the mating genes may have been inactivated in some strains. Despite this, phylogenetic analyses and linkage disequilibrium analyses indicate that H. werneckii is asexual.
(Text from Wikipedia and [image] (https://commons.wikimedia.org/wiki/File:Hortaea-werneckii-fungus--causes-tinea-nigra.jpg) from Wikipedia (http://en.wikipedia.org), the free encyclopaedia.
Assembly
The assembly presented has been imported from INSDC and has the assembly accession GCA_003704355.1.
Annotation
The annotation presented is derived from annotation submitted to INSDC with the assembly accession [GCA_003704355.1] (http://www.ebi.ac.uk/ena/data/view/GCA_003704355.1), with additional non-coding genes from Rfam. For more details, please visit INSDC annotation import.
More information
General information about this species can be found in Wikipedia.
Statistics
Summary
| Assembly | ASM370435v1, INSDC Assembly GCA_003704355.1, |
| Database version | 114.1 |
| Golden Path Length | 48,519,428 |
| Genebuild by | Biotechnical Faculty, University of Ljubljana |
| Genebuild method | Import |
| Data source | Biotechnical Faculty, University of Ljubljana |
Gene counts
| Coding genes | 16,503 |
| Gene transcripts | 16,503 |



 {#wiki_icon}
{#wiki_icon}![Follow us on Twitter! [twitter logo]](/i/twitter.png)
