elongin C
SuperContig supercont5.2: 640,478-642,038 reverse strand.
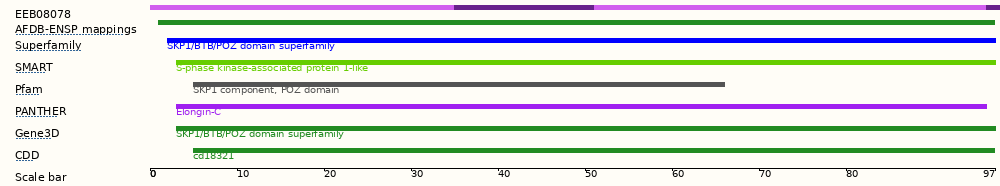
This transcript has 4 exons and is annotated with 7 domains and features.
This transcript is a product of gene SJAG_03211 Show transcript tableHide transcript table
elongin C
SuperContig supercont5.2: 640,478-642,038 reverse strand.
This transcript has 4 exons and is annotated with 7 domains and features.
This transcript is a product of gene SJAG_03211 Show transcript tableHide transcript table
.
Coiled-coil regions predicted by Ncoils.
URL to turn this track on
Copy the above url to force this track to be turned on
Click on the star to add/remove this track from your favourites
Click on the cross to turn the track off
Click to highlight/unhighlight this track
NCBIfam is a collection of protein families based on Hidden Markov Models (HMMs).
URL to turn this track on
Copy the above url to force this track to be turned on
Click on the star to add/remove this track from your favourites
Click on the cross to turn the track off
Click to highlight/unhighlight this track
Protein domains and motifs from the PROSITE patterns database.
URL to turn this track on
Copy the above url to force this track to be turned on
Click on the star to add/remove this track from your favourites
Click on the cross to turn the track off
Click to highlight/unhighlight this track
Signal peptide cleavage sites predicted by SignalP.
URL to turn this track on
Copy the above url to force this track to be turned on
Click on the star to add/remove this track from your favourites
Click on the cross to turn the track off
Click to highlight/unhighlight this track
Protein fingerprints (groups of conserved motifs) from the PRINTS database.
URL to turn this track on
Copy the above url to force this track to be turned on
Click on the star to add/remove this track from your favourites
Click on the cross to turn the track off
Click to highlight/unhighlight this track
Protein domains and motifs from the PIR (Protein Information Resource) Superfamily database.
URL to turn this track on
Copy the above url to force this track to be turned on
Click on the star to add/remove this track from your favourites
Click on the cross to turn the track off
Click to highlight/unhighlight this track
Protein domains and motifs from the PROSITE profiles database.
URL to turn this track on
Copy the above url to force this track to be turned on
Click on the star to add/remove this track from your favourites
Click on the cross to turn the track off
Click to highlight/unhighlight this track
Protein domains and motifs from the TIGRFAM database.
URL to turn this track on
Copy the above url to force this track to be turned on
Click on the star to add/remove this track from your favourites
Click on the cross to turn the track off
Click to highlight/unhighlight this track
Transmembrane helices predicted by TMHMM.
URL to turn this track on
Copy the above url to force this track to be turned on
Click on the star to add/remove this track from your favourites
Click on the cross to turn the track off
Click to highlight/unhighlight this track
Low complexity peptide sequences identified by Seg.
URL to turn this track on
Copy the above url to force this track to be turned on
Click on the star to add/remove this track from your favourites
Click on the cross to turn the track off
Click to highlight/unhighlight this track
Intrinsically disordered regions predicted by MobiDB lite.
URL to turn this track on
Copy the above url to force this track to be turned on
Click on the star to add/remove this track from your favourites
Click on the cross to turn the track off
Click to highlight/unhighlight this track
Structure-Function Linkage Database families.
URL to turn this track on
Copy the above url to force this track to be turned on
Click on the star to add/remove this track from your favourites
Click on the cross to turn the track off
Click to highlight/unhighlight this track
HAMAP families.
URL to turn this track on
Copy the above url to force this track to be turned on
Click on the star to add/remove this track from your favourites
Click on the cross to turn the track off
Click to highlight/unhighlight this track
Protein features that represent the mapping between Ensembl proteins (ENSP) and AlphaFoldDB protein structures (including their corresponding chains). Imported via UniProt mappings
URL to turn this track on
Copy the above url to force this track to be turned on
Click on the star to add/remove this track from your favourites
Click on the cross to turn the track off
Click to highlight/unhighlight this track
Protein domains and motifs from the SUPERFAMILY database.
URL to turn this track on
Copy the above url to force this track to be turned on
Click on the star to add/remove this track from your favourites
Click on the cross to turn the track off
Click to highlight/unhighlight this track
Protein domains and motifs from the SMART database.
URL to turn this track on
Copy the above url to force this track to be turned on
Click on the star to add/remove this track from your favourites
Click on the cross to turn the track off
Click to highlight/unhighlight this track
Protein domains and motifs from the Pfam database.
URL to turn this track on
Copy the above url to force this track to be turned on
Click on the star to add/remove this track from your favourites
Click on the cross to turn the track off
Click to highlight/unhighlight this track
PANTHER families.
URL to turn this track on
Copy the above url to force this track to be turned on
Click on the star to add/remove this track from your favourites
Click on the cross to turn the track off
Click to highlight/unhighlight this track
CATH/Gene3D families.
URL to turn this track on
Copy the above url to force this track to be turned on
Click on the star to add/remove this track from your favourites
Click on the cross to turn the track off
Click to highlight/unhighlight this track
Conserved Domain Database models.
URL to turn this track on
Copy the above url to force this track to be turned on
Click on the star to add/remove this track from your favourites
Click on the cross to turn the track off
Click to highlight/unhighlight this track
Ave. residue weight: 114.987 g/mol
Charge: -9.0
Isoelectric point: 4.1310
Molecular weight: 11,153.76 g/mol
Number of residues: 97 aa
.