Aspergillus niger Assembly and Gene Annotation
About the Aspergillus niger genome
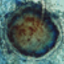
Aspergillus niger causes black mould on certain fruits and vegetables such as grapes, apricots, onions, and peanuts, and is a common contaminant of food. It is ubiquitous in soil and is commonly reported from indoor environments.
Picture credit: Aspergillus/Aspergillosis Website owned by the Fungal Research Trust
The Department of Energy (DOE) Joint Genome Institute (JGI)](http://genome.jgi-psf.org/Aspni1/Aspni1.home.html) sequenced the genome of Aspergillus niger strain ATCC 1015 at 8.9X coverage using whole genome shotgun (WGS) sequencing. The version 3.0 of the assembly (June 30, 2008) contains 8 chromosomes, made up of 21 scaffolds, and 3 unmapped scaffolds. The 7 of the 8 chromosomes and 1 unmapped scaffold contain telomeres making A. niger's genome 34.85 MB in size.
References
- CADRE: the Central Aspergillus Data
REpository.
Mabey JE, Anderson MJ, Giles PF, Miller CJ, Attwood TK, Paton NW, Bornberg-Bauer E, Robson GD, Oliver SG, Denning DW. 2004. Nucleic Acids Res.. 32:D401-5. - Genome sequencing and analysis of the versatile cell factory
Aspergillus niger CBS
513.88.
Pel HJ, de Winde JH, Archer DB, Dyer PS, Hofmann G, Schaap PJ, Turner G, de Vries RP, ..., van Ooyen AJ et al. 2007. Nat. Biotechnol.. 25:221-231.
More information
General information about this species can be found in Wikipedia.
Statistics
Summary
| Assembly | ASM285v2, INSDC Assembly GCA_000002855.2, Mar 2015 |
| Database version | 114.2 |
| Golden Path Length | 33,975,768 |
| Genebuild by | |
| Genebuild method | Import |
| Data source | CADRE |
Gene counts
| Coding genes | 13,942 |
| Non coding genes | 679 |
| Small non coding genes | 679 |
| Pseudogenes | 151 |
| Gene transcripts | 14,772 |



![Follow us on Twitter! [twitter logo]](/i/twitter.png)
