BioMart RESTful access (Perl and wget)
BioMart RESTful access is a quick and easy way to query the Ensembl marts using wget or perl and doesn't require any programing knowledge.
Summary
- Obtaining the BioMart xml from the BioMart website
- Using the wget UNIX command
- Using the BioMart PERL API script
- The xml Completion Stamp
Obtaining the BioMart xml from the BioMart website
You can easily obtain a BioMart xml file from the BioMart interface. For example, navigate to the Ensembl gene mart on the Ensembl website, apply your required filters and select the attribute you are insterested in. As shown in the example below, filter on the human Ensembl Gene ID "ENSG00000139618" and select the Ensembl Gene, Transcript IDs, HGNC symbols and Uniprot Swissprot accessions attribute. The BioMart xml file can be downloaded from the BioMart result page accessible via the "Results" button. To get your BioMart query in xml, just click on the xml button as indicated by the red box in the image below.
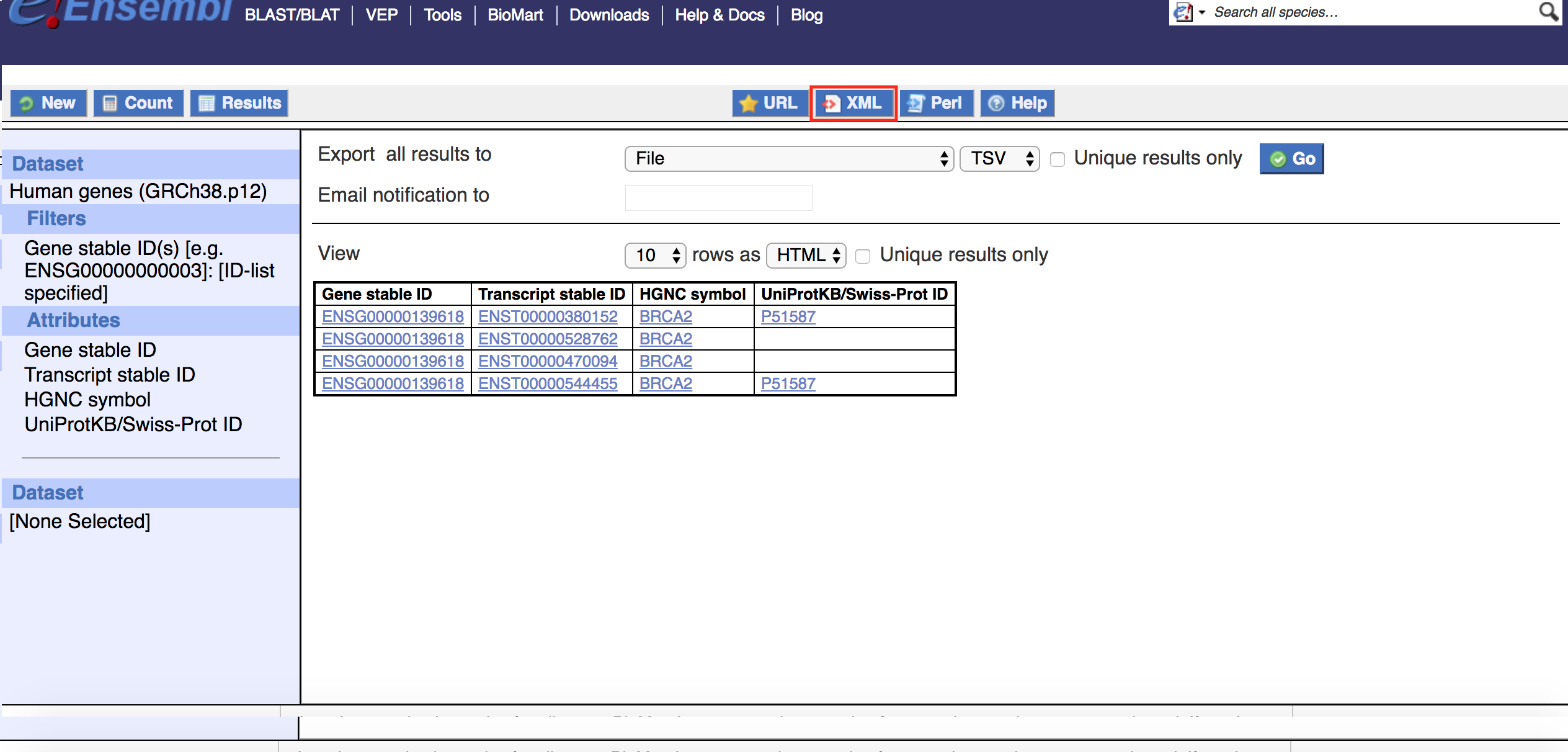
The xml button will open a new browser window and display the BioMart query in xml format, the text will be similar to the following image.
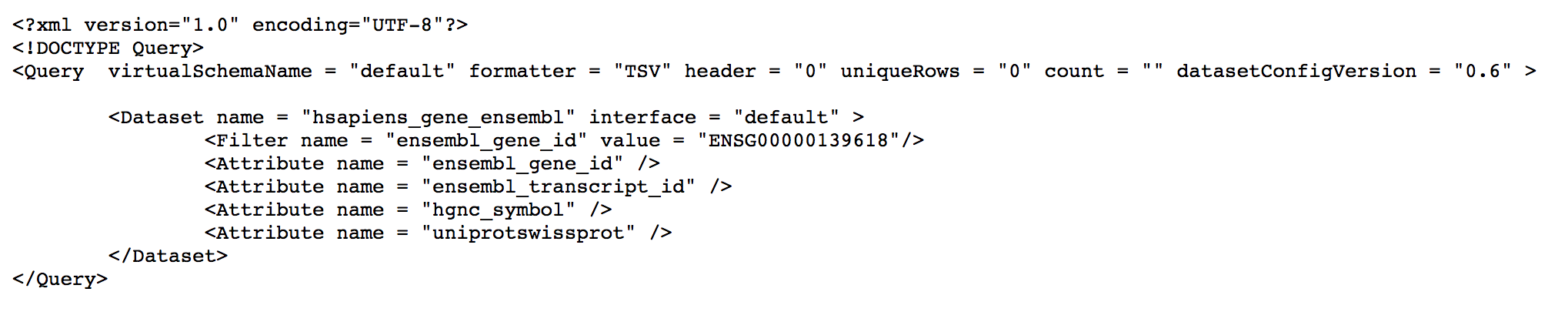
Just save the content of this page in a new file on your computer, e.g 'hgnc_swissprot.xml' in our example.
Using the wget UNIX command
Type the following command in your terminal:
wget -O result.txt 'https://www.ensembl.org/biomart/martservice?query=
Then copy the content of the previouly saved xml file all in one line after the "query=", you should now have the following:
wget -O result.txt 'https://www.ensembl.org/biomart/martservice?query=<?xml version="1.0" encoding="UTF-8"?><!DOCTYPE Query><Query virtualSchemaName = "default" formatter = "TSV" header = "0" uniqueRows = "0" count = "" datasetConfigVersion = "0.6" ><Dataset name = "hsapiens_gene_ensembl" interface = "default" ><Filter name = "ensembl_gene_id" value = "ENSG00000139618"/><Attribute name = "ensembl_gene_id" /><Attribute name = "ensembl_transcript_id" /><Attribute name = "hgnc_symbol" /><Attribute name = "uniprotswissprot" /></Dataset></Query>'
Finally, just run the command to get the BioMart data stored inside the "result.txt" file. In our example, we get the following result.txt file:
less result.txt ENSG00000139618 ENST00000380152 BRCA2 P51587 ENSG00000139618 ENST00000528762 BRCA2 ENSG00000139618 ENST00000470094 BRCA2 ENSG00000139618 ENST00000544455 BRCA2 P51587
Using the BioMart PERL API script
First, you will need to download the BioMart API, to do this you can follow the command below:
git clone --branch cvs/release-0_7 https://github.com/biomart/biomart-perl
To use the Ensembl marts from the ensembl.org website, just edit the path variable in the biomart-perl/scripts/webExample.pl Perl script to the following:
my $path="https://www.ensembl.org/biomart/martservice?";
Finally run the biomart-perl/scripts/webExample.pl Perl script with the xml file obtained in the "Obtaining the BioMart xml" section:
biomart-perl/scripts: perl webExample.pl hgnc_swissprot.xml
You should get an output similar to the following:
ENSG00000139618 ENST00000380152 BRCA2 P51587 ENSG00000139618 ENST00000528762 BRCA2 ENSG00000139618 ENST00000470094 BRCA2 ENSG00000139618 ENST00000544455 BRCA2 P51587
The xml Completion Stamp
If you want to make sure you are getting all the data from your BioMart query, you can add a "CompletionStamp" to the xml file. To do this, just open the previously obtained xml file in the "Obtaining the BioMart xml" section and add the following text in the query tag:
completionStamp = "1"
The above command should be paste in the location indicated by the red box in the image below:

Then either use the wget command or the BioMart Perl script. If the query successfuly ran, you will get a "[success]" after running the wget or BioMart perl script:
biomart-perl/scripts: perl webExample.pl hgnc_swissprot_completionstamp.xml ENSG00000139618 ENST00000380152 BRCA2 P51587 ENSG00000139618 ENST00000528762 BRCA2 ENSG00000139618 ENST00000470094 BRCA2 ENSG00000139618 ENST00000544455 BRCA2 P51587 [success]




![Follow us on Twitter! [twitter logo]](/i/twitter.png)
